Author: Nick Semenkovich (semenko@alum.mit.edu)
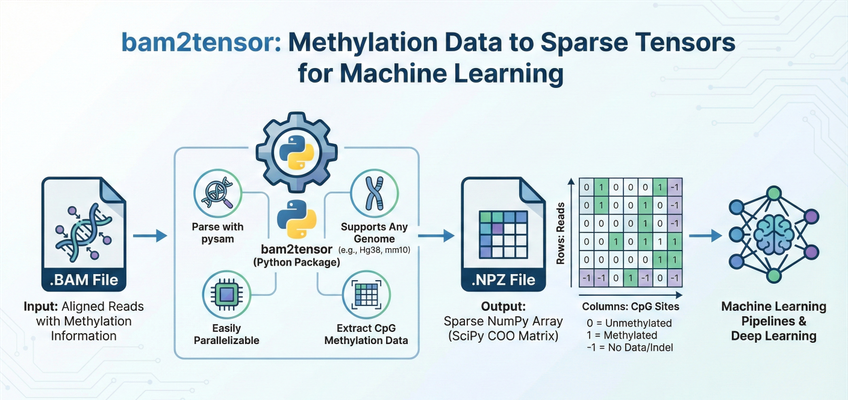
bam2tensor is a Python package for converting bisulfite-sequencing (BS-seq) and enzymatic methylation sequencing (EM-seq) .bam files to sparse tensor representations of DNA methylation data. It extracts read-level methylation states from CpG sites and outputs efficient sparse COO matrices as .npz files, ready for deep learning pipelines.
- Features
- Requirements
- Installation
- Quick Start
- Usage
- Filtering Conversion Errors
- Inspecting Output Files
- Output Data Structure
- Supported Aligners
- Performance Tips
- API Reference
- Contributing
- AI Coding Agents
- License
- Credits
- BAM Parsing: Efficiently parses
.bamfiles using pysam - Complete CpG Extraction: Extracts methylation data from all CpG sites genome-wide
- Multi-Genome Support: Works with any reference genome (GRCh38/hg38, T2T-CHM13, mm10, etc.)
- Sparse Storage: Stores data in sparse COO matrix format for memory-efficient loading
- NumPy/SciPy Integration: Exports to
.npzfiles compatible with NumPy and SciPy - Efficient Algorithm: Linear-scan algorithm ensures minimal memory usage with no read duplication
- Batch Processing: Process multiple BAM files with directory recursion
- Caching: CpG site indexing is cached to accelerate repeated runs on the same genome
- Quality Filtering: Configurable mapping quality thresholds
- Per-Read Fragment Length: Stores BAM TLEN (template length) alongside the methylation tensor for joint fragment-methylation analysis
- Conversion-Error Filters: Optional per-read filters for incomplete bisulfite/EM-seq conversion (ported from
nebiolabs/mark-nonconverted-reads) and EM-seq fragment-level over-conversion (Loyfer et al. 2026)
- Python 3.10 or higher
- A reference genome FASTA file (must match the genome used for alignment)
- Indexed BAM files (
.bamwith corresponding.bam.baiindex files)
Core dependencies are automatically installed:
pysam- BAM file handlingbiopython- FASTA parsingscipy- Sparse matrix operationsnumpy- Numerical operationsclick- Command-line interfacetqdm- Progress bars
pip install bam2tensorgit clone https://github.com/mcwdsi/bam2tensor.git
cd bam2tensor
pip install .git clone https://github.com/mcwdsi/bam2tensor.git
cd bam2tensor
uv sync# Basic usage with a single BAM file
bam2tensor \
--input-path sample.bam \
--reference-fasta GRCh38.fa \
--genome-name hg38
# This creates: sample.methylation.npzProcess a single bisulfite-sequencing or EM-seq BAM file:
bam2tensor \
--input-path /path/to/aligned_reads.bam \
--reference-fasta /path/to/reference.fa \
--genome-name hg38This will:
- Parse the reference FASTA to identify all CpG sites (cached for future runs)
- Extract methylation states from each read in the BAM file
- Output a sparse matrix to
aligned_reads.methylation.npz
Don't have a local reference FASTA? bam2tensor can download and cache common reference genomes automatically:
# Download hg38 and process a BAM file in one command
bam2tensor \
--input-path sample.bam \
--download-reference hg38
# List available genomes
bam2tensor --input-path sample.bam --list-genomesAvailable genomes: hg38, hg19, mm10, T2T-CHM13. Downloaded references are cached in ~/.cache/bam2tensor/ for future runs.
Process all BAM files in a directory recursively:
bam2tensor \
--input-path /path/to/bam_directory/ \
--reference-fasta /path/to/reference.fa \
--genome-name hg38 \
--verboseEach BAM file will generate a corresponding .methylation.npz file in the same location.
Write output files to a specific directory instead of next to the input BAMs:
bam2tensor \
--input-path /shared/bams/ \
--reference-fasta /shared/ref/GRCh38.fa \
--genome-name hg38 \
--output-dir /scratch/methylation_output/This is useful on HPC clusters where input directories may be read-only or on slow shared storage.
For non-human genomes or custom chromosome sets:
# Mouse genome (mm10)
bam2tensor \
--input-path mouse_sample.bam \
--reference-fasta mm10.fa \
--genome-name mm10 \
--expected-chromosomes "chr1,chr2,chr3,chr4,chr5,chr6,chr7,chr8,chr9,chr10,chr11,chr12,chr13,chr14,chr15,chr16,chr17,chr18,chr19,chrX,chrY"
# T2T-CHM13 human genome
bam2tensor \
--input-path sample.bam \
--reference-fasta chm13v2.0.fa \
--genome-name T2T-CHM13 \
--expected-chromosomes "chr1,chr2,chr3,chr4,chr5,chr6,chr7,chr8,chr9,chr10,chr11,chr12,chr13,chr14,chr15,chr16,chr17,chr18,chr19,chr20,chr21,chr22,chrX,chrY"Usage: bam2tensor [OPTIONS]
Extract read-level methylation data from an aligned bisulfite-seq or EM-seq
.bam file and export the data as a SciPy sparse matrix.
Options:
--version Show the version and exit.
--input-path PATH Input .bam file OR directory to recursively
process. [required]
--genome-name TEXT A custom string referring to your genome
name, used to save a cache file (e.g. hg38,
hg39-no-alt, etc.).
--expected-chromosomes TEXT A comma-separated list of chromosomes to
expect in the .fa genome. Defaults to hg38
chromosomes.
--reference-fasta FILE Reference genome fasta file (critical to
determine CpG sites).
--quality-limit INTEGER Quality filter for aligned reads (default =
20)
--filter-non-converted Drop reads with >= --non-converted-threshold
retained non-CpG cytosines, the signature of
incomplete bisulfite/EM-seq conversion (port
of nebiolabs/mark-nonconverted-reads).
Default: off.
--non-converted-threshold INTEGER
Minimum count of retained non-CpG cytosines
to drop a read (default = 3, matches NEB
mark-nonconverted-reads).
--filter-em-overconversion Drop EM-seq reads whose covered CpGs are all
called unmethylated and cover at least --em-
overconversion-min-cpgs sites (heuristic for
the fragment-level over-conversion artifact
described in Loyfer et al. bioRxiv
2026.03.24.713040). Default: off.
--em-overconversion-min-cpgs INTEGER
Minimum covered CpG count required before
the EM over-conversion filter will drop a
read (default = 3).
--verbose Verbose output.
--skip-cache De-novo generate CpG sites (slow).
--debug Debug mode (extensive validity checking +
debug messages).
--overwrite Overwrite output file if it exists.
--output-dir DIRECTORY Output directory for .methylation.npz files.
Defaults to same directory as the input BAM.
--download-reference [t2t-chm13|hg19|hg38|mm10]
Download and cache a known reference genome
(e.g. hg38, hg19, mm10, T2T-CHM13). Use
--list-genomes to see options.
--list-genomes List available reference genomes for
download and exit.
--help Show this message and exit.
| Option | Description |
|---|---|
--input-path |
Path to a single .bam file or a directory. If a directory is provided, all .bam files are processed recursively. |
--genome-name |
An identifier for your reference genome (e.g., hg38, mm10). Used to name the cache file for CpG site positions. |
--expected-chromosomes |
Comma-separated list of chromosome names to process. Chromosomes not in this list are skipped. Defaults to human autosomes + sex chromosomes. |
--reference-fasta |
Path to the reference genome FASTA file. Must match the genome used for alignment. |
--quality-limit |
Minimum mapping quality score (MAPQ) for reads to be included. Default is 20. |
--filter-non-converted |
Drop reads with retained non-CpG cytosines above --non-converted-threshold (incomplete conversion). See Filtering Conversion Errors. |
--non-converted-threshold |
Threshold for the non-converted filter. Default is 3. |
--filter-em-overconversion |
Drop EM-seq reads whose covered CpGs are all unmethylated and cover ≥ --em-overconversion-min-cpgs sites. See Filtering Conversion Errors. |
--em-overconversion-min-cpgs |
Minimum covered CpG count before the EM over-conversion filter will drop a read. Default is 3. |
--verbose |
Enable detailed progress output including per-chromosome progress bars. |
--skip-cache |
Force regeneration of CpG site cache. Useful if you've modified the reference or chromosome list. |
--debug |
Enable extensive validation and debug output. Slower but useful for troubleshooting. |
--overwrite |
Overwrite existing .methylation.npz files. Without this flag, existing outputs are skipped. |
--output-dir |
Write .methylation.npz files to this directory instead of next to the input BAMs. Created automatically if it doesn't exist. |
--download-reference |
Download and cache a known reference genome. Choices: hg38, hg19, mm10, T2T-CHM13. Replaces --reference-fasta. |
--list-genomes |
List available reference genomes for --download-reference and exit. |
Bisulfite and EM-seq library preparation can produce two kinds of per-read conversion errors that bias downstream methylation calls. bam2tensor provides two opt-in filters to drop affected reads at extraction time. Both are default-off, apply per read, and are recorded in the output metadata.json so downstream consumers know which filters were applied.
Ports the logic of nebiolabs/mark-nonconverted-reads. A read is dropped if it carries at least --non-converted-threshold (default 3) retained non-CpG cytosines, a signature of incomplete bisulfite or EM-seq conversion.
- Bismark BAMs: counted directly from the
XMtag's uppercaseH/X/Ucharacters (retained cytosines in CHH/CHG/unknown contexts). - Biscuit / bwameth / gem3 BAMs: counted by comparing the read to the reference via the
MDtag (using pysam'sget_aligned_pairs(with_seq=True)). SNPs — where the read's retainedCsits over a reference base that isn'tC— are excluded from the count, matching NEB's reference-validation step. No separate FASTA reload is required.
A heuristic inspired by Loyfer et al. (bioRxiv 2026.03.24.713040). That paper shows EM-seq reproducibly produces ~1–2.5% of multi-CpG fragments that appear fully unmethylated across every covered CpG — a fragment-level artifact absent from WGBS and Oxford Nanopore. This filter drops any read whose covered CpGs are all called unmethylated and cover at least --em-overconversion-min-cpgs sites (default 3, the regime where the EM-seq artifact is clearly separable from WGBS in Loyfer et al. Fig. 1C).
The filter is a blunt instrument: it will also drop genuinely fully-unmethylated biological fragments at unmethylated markers. Enable it only when your downstream application (e.g., cfDNA deconvolution at constitutively methylated loci) can tolerate that trade-off.
bam2tensor \
--input-path sample.bam \
--reference-fasta GRCh38.fa \
--genome-name hg38 \
--filter-non-converted \
--filter-em-overconversionFilter parameters and enabled state are written to the output metadata.json:
{
"filters": {
"non_converted_reads": {"enabled": true, "threshold": 3},
"em_overconversion": {"enabled": true, "min_cpgs": 3}
}
}The two filters differ in whether they can be replayed downstream without the source BAM:
-
--filter-em-overconversionis reproducible from the.npzalone. The heuristic is a pure function of each row's CpG state values. A downstream consumer who receives an unfiltered.npzcan replay the filter at analysis time:import scipy.sparse mat = scipy.sparse.load_npz("sample.methylation.npz").tocsr() min_cpgs = 3 kept_rows = [] for i in range(mat.shape[0]): row = mat.getrow(i).toarray().ravel() covered = row[(row == 0) | (row == 1)] # drop -1 no-data is_overconv = len(covered) >= min_cpgs and (covered == 0).all() if not is_overconv: kept_rows.append(i)
-
--filter-non-convertedis not reproducible from the.npzalone. It relies on retained non-CpG cytosines (or Bismark'sH/X/U), which are never written to the matrix. If you need this filter, apply it at extraction time (or re-run bam2tensor against the original BAM).
Use bam2tensor-inspect to view a summary of any .methylation.npz file without writing Python:
$ bam2tensor-inspect sample.methylation.npz
sample.methylation.npz
Genome: hg38
Chromosomes: 24 (chr1, chr2, ... chrX, chrY)
Reads: 1,423,891
CpG sites: 28,217,448
Data points: 12,847,322 (sparsity: 99.97%)
Fragment len: median 167, mean 182, range [50, 600]
Filters: non-converted (>= 3 non-CpG Cs)
EM over-conversion (all-unmethylated, >= 3 CpGs)
CpG index CRC32: a1b2c3d4
bam2tensor: v2.5
File size: 14.2 MBWhen no filters were applied, the line reads Filters: none. Files produced by bam2tensor versions older than v2.5 omit the line entirely.
You can pass multiple files at once:
$ bam2tensor-inspect *.methylation.npzThis works on files produced by older versions of bam2tensor too (metadata fields will be omitted).
bam2tensor generates one .methylation.npz file per input BAM file. Each file contains a SciPy sparse COO matrix (scipy.sparse.coo_matrix) with the following structure:
| Dimension | Represents |
|---|---|
| Rows | Unique sequencing reads (primary alignments that pass quality and flag filters, numbered sequentially as encountered across chromosomes) |
| Columns | CpG sites from the reference genome, ordered by genomic position across all chromosomes (chr1, chr2, ..., chrX, chrY). Column i maps to the i-th CpG dinucleotide in the reference FASTA. |
The column dimension is determined entirely by the reference genome: it equals the total number of CpG sites across all --expected-chromosomes. For example, hg38 with default chromosomes has ~28 million CpG columns. To map column indices back to genomic coordinates (e.g., column 12345 → chr1:29503), use the GenomeMethylationEmbedding class with the same reference FASTA and chromosome list (see Working with Genomic Coordinates below).
| Value | Meaning |
|---|---|
1 |
Methylated (cytosine preserved as C after bisulfite/enzymatic conversion) |
0 |
Unmethylated (cytosine converted to T by bisulfite/enzymatic treatment) |
-1 |
No data (indel, SNV, or other non-C/T base at a CpG position) |
Note: The matrix uses SciPy's COO sparse format, which explicitly stores all non-zero values. Unmethylated sites (value 0) are stored as explicit entries. Positions not covered by a read are simply absent from the matrix (implicit zero, which is distinct from the explicit 0 = unmethylated).
Each .methylation.npz file includes a tlen.npy entry inside the ZIP archive containing the signed BAM template length (TLEN) for every read in the matrix. This enables joint fragment-length and methylation analysis without re-processing the BAM.
- One
int32value per read (row), in the same order as the sparse matrix rows - Signed: positive for the leftmost read in a pair, negative for the rightmost
- Zero for single-end reads or reads with unmapped mates
- Use
abs(tlen)to get fragment lengths
from bam2tensor.metadata import read_npz_tlen
import numpy as np
tlen = read_npz_tlen("sample.methylation.npz")
if tlen is not None:
frag_lengths = np.abs(tlen)
nonzero = frag_lengths[frag_lengths > 0]
print(f"Median fragment length: {np.median(nonzero):.0f}")
print(f"Mean fragment length: {np.mean(nonzero):.0f}")Each .methylation.npz file includes a metadata.json entry inside the ZIP archive with provenance information:
| Field | Description |
|---|---|
bam2tensor_version |
Version of bam2tensor that produced the file |
genome_name |
Genome identifier (e.g., hg38, mm10) |
expected_chromosomes |
List of chromosomes included in the column mapping |
total_cpg_sites |
Total number of CpG columns in the matrix |
cpg_index_crc32 |
CRC32 checksum of the CpG site positions (verifies identical column semantics) |
filters |
Nested dict recording which opt-in conversion-error filters were applied (non_converted_reads, em_overconversion) and their parameters. See Filtering Conversion Errors. Added in v2.5. |
This metadata is ignored by scipy.sparse.load_npz, so existing code continues to work. To read it:
from bam2tensor.metadata import read_npz_metadata
meta = read_npz_metadata("sample.methylation.npz")
if meta is not None:
print(f"Genome: {meta['genome_name']}")
print(f"CpG sites: {meta['total_cpg_sites']:,}")
print(f"CpG index CRC32: {meta['cpg_index_crc32']}")The cpg_index_crc32 field uniquely identifies the column mapping. Two files with the same CRC32 have identical column semantics (same chromosomes, same CpG positions, same order) and their matrices can be directly stacked or compared. The metadata is also accessible without bam2tensor installed, since .npz files are ZIP archives:
unzip -p sample.methylation.npz metadata.json | python -m json.toolimport scipy.sparse
import numpy as np
# Load the sparse matrix
methylation_matrix = scipy.sparse.load_npz("sample.methylation.npz")
print(f"Matrix shape: {methylation_matrix.shape}")
print(f"Number of reads: {methylation_matrix.shape[0]}")
print(f"Number of CpG sites: {methylation_matrix.shape[1]}")
print(f"Non-zero entries: {methylation_matrix.nnz}")
print(f"Sparsity: {1 - methylation_matrix.nnz / np.prod(methylation_matrix.shape):.4%}")For small regions or when dense operations are needed:
# Convert entire matrix to dense (warning: may use significant memory)
dense_matrix = methylation_matrix.toarray()
# Convert to CSR format for efficient row slicing
csr_matrix = methylation_matrix.tocsr()
# Get methylation data for reads 0-99
subset = csr_matrix[0:100, :].toarray()
# Convert to CSC format for efficient column slicing
csc_matrix = methylation_matrix.tocsc()
# Get data for CpG sites 1000-1099
cpg_subset = csc_matrix[:, 1000:1100].toarray()To map between matrix column indices and genomic coordinates, use the GenomeMethylationEmbedding class:
from bam2tensor.embedding import GenomeMethylationEmbedding
# Load or recreate the embedding used during extraction
embedding = GenomeMethylationEmbedding(
genome_name="hg38",
expected_chromosomes=["chr" + str(i) for i in range(1, 23)] + ["chrX", "chrY"],
fasta_source="/path/to/GRCh38.fa",
)
# Convert matrix column index to genomic position
chrom, pos = embedding.embedding_to_genomic_position(12345)
print(f"Column 12345 corresponds to {chrom}:{pos}")
# Convert genomic position to matrix column index
col_idx = embedding.genomic_position_to_embedding("chr1", 10525)
print(f"chr1:10525 is at column {col_idx}")
# Get total number of CpG sites
print(f"Total CpG sites: {embedding.total_cpg_sites:,}")import scipy.sparse
import numpy as np
# Load the data
matrix = scipy.sparse.load_npz("sample.methylation.npz")
csr = matrix.tocsr()
# Calculate per-CpG methylation rates (excluding -1 values)
methylation_rates = []
for cpg_idx in range(matrix.shape[1]):
col_data = csr.getcol(cpg_idx).toarray().flatten()
# Filter out -1 (no data) and positions with no coverage
valid_data = col_data[(col_data >= 0)]
if len(valid_data) > 0:
rate = np.mean(valid_data)
else:
rate = np.nan
methylation_rates.append(rate)
methylation_rates = np.array(methylation_rates)
print(f"Mean methylation rate: {np.nanmean(methylation_rates):.2%}")
print(f"CpG sites with coverage: {np.sum(~np.isnan(methylation_rates)):,}")import torch
import scipy.sparse
import numpy as np
# Load sparse matrix
matrix = scipy.sparse.load_npz("sample.methylation.npz")
# Convert to PyTorch sparse tensor
coo = matrix.tocoo()
indices = torch.LongTensor(np.vstack((coo.row, coo.col)))
values = torch.FloatTensor(coo.data)
shape = torch.Size(coo.shape)
sparse_tensor = torch.sparse_coo_tensor(indices, values, shape)
print(f"PyTorch sparse tensor shape: {sparse_tensor.shape}")
# For models that need dense input (specific region)
region_start, region_end = 0, 1000
dense_region = matrix.tocsc()[:, region_start:region_end].toarray()
dense_tensor = torch.FloatTensor(dense_region)bam2tensor supports BAM files from bisulfite-aware and EM-seq-aware aligners that include strand information tags:
| Aligner | Tag | Values |
|---|---|---|
| Bismark | XM |
Z (methylated CpG), z (unmethylated CpG) |
| Biscuit | YD |
f (forward/OT/CTOT), r (reverse/OB/CTOB) |
| bwameth | YD |
f (forward/OT/CTOT), r (reverse/OB/CTOB) |
| gem3 / Blueprint | XB |
C (forward), G (reverse) |
These tags indicate which strand the original bisulfite/EM-seq-converted DNA came from, which is essential for correctly interpreting C/T as methylated/unmethylated.
Note: EM-seq (enzymatic methyl-seq) data produces the same C-to-T conversion pattern as bisulfite sequencing and is fully supported when aligned with any of the above tools.
-
Use the cache: The first run on a new genome builds a CpG site index, which is cached. Subsequent runs are much faster.
-
Process in parallel: bam2tensor processes one BAM at a time, but you can run multiple instances in parallel on different BAM files:
# Using GNU parallel find /data/bams -name "*.bam" | parallel -j 4 \ bam2tensor --input-path {} --reference-fasta ref.fa --genome-name hg38
-
Ensure BAM files are indexed: Each BAM file requires a corresponding
.bam.baiindex file. Create with:samtools index sample.bam
-
Use SSDs: Both reading BAM files and writing output benefit from fast storage.
-
Memory considerations: Memory usage scales with the number of CpG sites (columns) rather than reads. For human genomes (~28M CpG sites), expect moderate memory usage.
Main class for managing CpG site positions and coordinate conversions.
GenomeMethylationEmbedding(
genome_name: str, # Identifier for caching
expected_chromosomes: list, # List of chromosome names to process
fasta_source: str, # Path to reference FASTA
skip_cache: bool = False, # Force regeneration of cache
verbose: bool = False # Enable verbose output
)Key Methods:
embedding_to_genomic_position(embedding: int) -> tuple[str, int]- Convert column index to (chromosome, position)genomic_position_to_embedding(chrom: str, pos: int) -> int- Convert genomic position to column index
Key Attributes:
total_cpg_sites: int- Total number of CpG sites across all chromosomescpg_sites_dict: dict[str, list[int]]- Dictionary mapping chromosome names to lists of CpG positions
Core function for extracting methylation data from a BAM file.
extract_methylation_data_from_bam(
input_bam: str, # Path to BAM file
genome_methylation_embedding: GenomeMethylationEmbedding, # Embedding object
quality_limit: int = 20, # Minimum MAPQ
filter_non_converted: bool = False, # Drop reads with retained non-CpG Cs
non_converted_threshold: int = 3, # Threshold for the above filter
filter_em_overconversion: bool = False, # Drop EM-seq fragment-level over-conversion reads
em_overconversion_min_cpgs: int = 3, # Min CpGs before applying the above filter
verbose: bool = False, # Enable verbose output
debug: bool = False # Enable debug output
) -> ExtractionResultReturns: An ExtractionResult named tuple with two fields:
matrix: A SciPy COO sparse matrix with shape (n_reads, n_cpg_sites)tlen: A 1-D numpyint32array of shape (n_reads,) containing the signed template length (BAM TLEN field) for each read
Read per-read template lengths from a .methylation.npz file.
read_npz_tlen(npz_path: str) -> np.ndarray | NoneReturns: The per-read template-length array, or None if the file was produced by an older version of bam2tensor.
Contributions are welcome! Please see the Contributor Guide for guidelines on:
- Setting up a development environment
- Running tests
- Code style requirements
- Submitting pull requests
# Install development dependencies
uv sync
# Run all checks (linting, type checking, tests)
nox
# Run specific checks
nox --session=tests # Run pytest
nox --session=mypy # Type checking
nox --session=pre-commit # Linting
# Format code
uv run black src tests
uv run ruff check --fix src testsThis project includes a CLAUDE.md file with detailed guidance for AI coding agents (Claude Code, Copilot, Cursor, etc.). If you're using an AI assistant to contribute, please ensure it follows the project conventions described there.
Key points for AI agents:
- Run
noxbefore committing to validate all checks pass - Follow Google-style docstrings validated by darglint (long strictness)
- Use
uv syncfor dependency management - See
CLAUDE.mdfor complete guidelines
Distributed under the terms of the MIT license, bam2tensor is free and open source software.
If you encounter any problems, please file an issue with:
- A description of the problem
- Steps to reproduce
- Your Python version and operating system
- Relevant error messages or logs
Created and maintained by Nick Semenkovich (@semenko), as part of the Medical College of Wisconsin's Data Science Institute.
This project was generated from Statistics Norway's SSB PyPI Template.